John Conway's Game of Life
Summer 2020 ~ @TitouanCh
Discover the Game of Life from the famous mathematician John Conway. A cellular automaton in your browser.
The Game of Life is easily one of if not the most famous cellular automaton. Despite it's simple rules, it can generates some remarquables structures.
John Conway
John Conway (1937-2020) was an English mathematician from Liverpool.
In October 1970, Scientific American published a column featuring some of his work, most notably Conway's Game of Life, a mathematical game that generates structures and patterns resembling real-life phenomena.
Conway was once quoted as saying he hated the Game of Life because he felt it overshadowed much of his other work, and that after solving the game he found it uninteresting. However, later in his life, he retracted many of these statements, expressing reconciliation with the game and stating that he was quite proud of it.
 Mathematician John Horton Conway
Mathematician John Horton Conway
Game of life and cellular automata
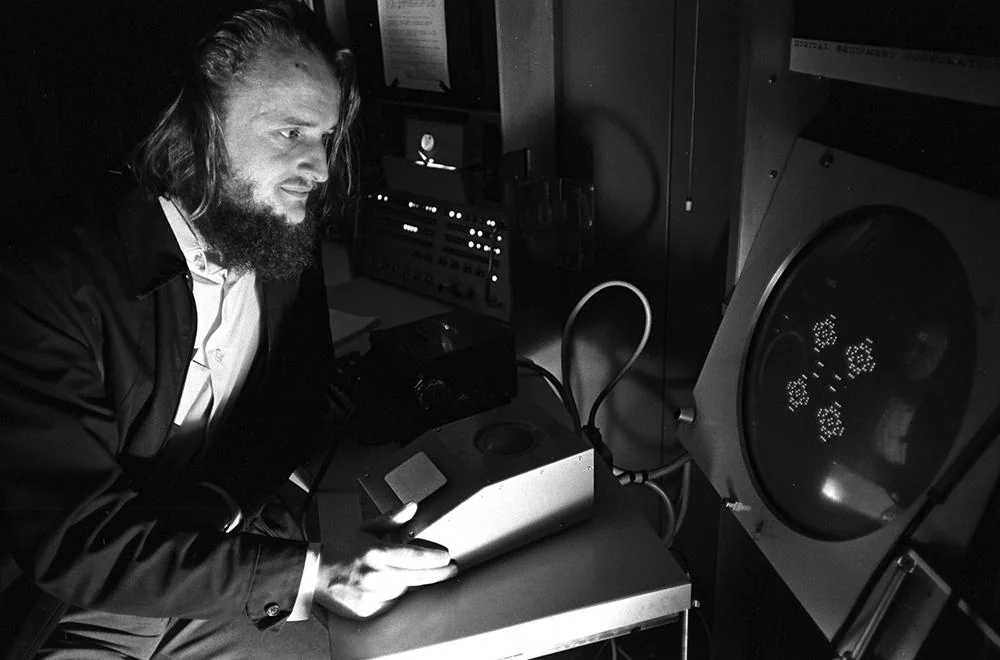

Conway playing Game of Life on a computer in 1970
John Conway (1937-2020) was an English mathematician from Liverpool.
In October 1970, Scientific American published a column featuring some of his work, most notably Conway's Game of Life, a mathematical game that generates structures and patterns resembling real-life phenomena.
Conway was once quoted as saying he hated the Game of Life because he felt it overshadowed much of his other work, and that after solving the game he found it uninteresting. However, later in his life, he retracted many of these statements, expressing reconciliation with the game and stating that he was quite proud of it.
 Mathematician John Horton Conway
Mathematician John Horton Conway
Conway playing Game of Life on a computer in 1970
The game of life is a cellular automaton. This means that the game is played on a grid of cells. At any point in time cells can be either dead or alive.
The game is not really a game in the sense that there is only one player. Cells evolve collectively across the grid, moving, appearing, and disappearing in each step according to three seemingly straightforward rules:
- An alive cell with 2 or 3 alive neighbors, stays alive.
- A dead cell with 3 alive neighbors, is colonised (becomes alive).
- All other cells stay dead or die.
My implementation
There are various approaches to implementing the Game of Life, but they generally fall into two main categories:
- CPU-bound simulations: The simplest method to implement the game of life is to create a grid of cells represented as a 2D array in CPU memory. The simulation progresses step by step by calculating the next state of each cell sequentially. This straightforward approach, though easy to understand, tends to be slow, especially as the grid size increases. In fact, its performance deteriorates exponentially with larger grids. While multi-threading in conjunction with space partitioning can be applied to improve speed, there are more efficient alternatives.
- GPU-bound simulations: A faster approach is to use the Graphics Processing Unit (GPU) to parallelize the Game of Life. The nature of the game makes it a perfect fit for GPU parallelization. GPUs are designed to handle lots of calculations simultaneously, which is exactly what is needed for updating thousands of cells at once.
For my implementation, I decided to utilize OpenGL. Implementation the game of life into two fragment shaders. One that simply copies the current state to an image buffer and a second that applies the game rules to the buffer. This approach is very fast, and can be played straight into the browser using WebGL as seen above.
Using OpenGL and shaders not only makes the Game of Life run smoothly but also shows off how powerful GPUs can be for handling simple simulations like this. It's a cool way to quickly dive into both graphics programming, GPU parralelization and the world of cellular automata which was my goal for this project.